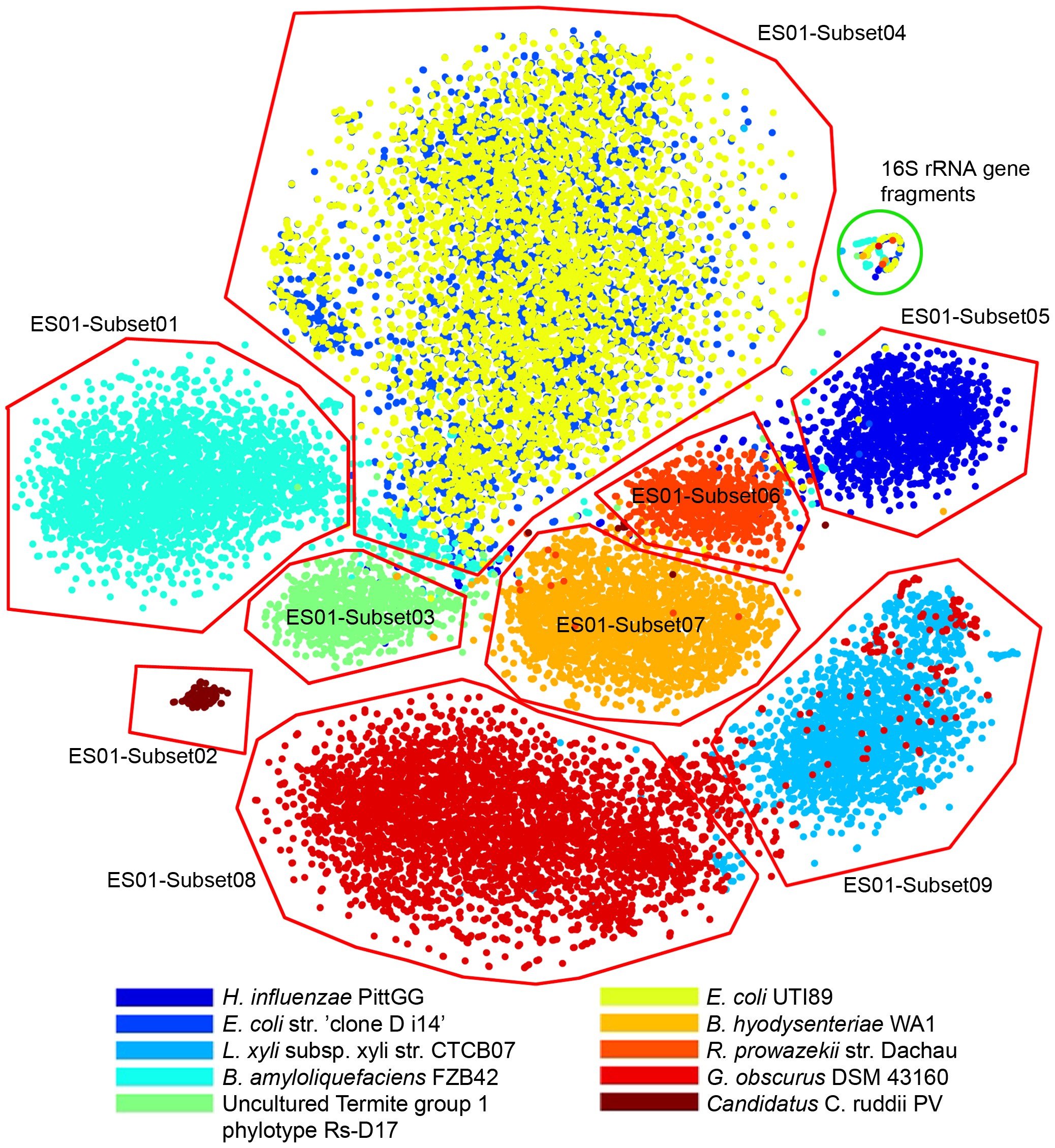
Alignment-free Visualization of Metagenomic Data by Nonlinear Dimension Reduction | Scientific Reports
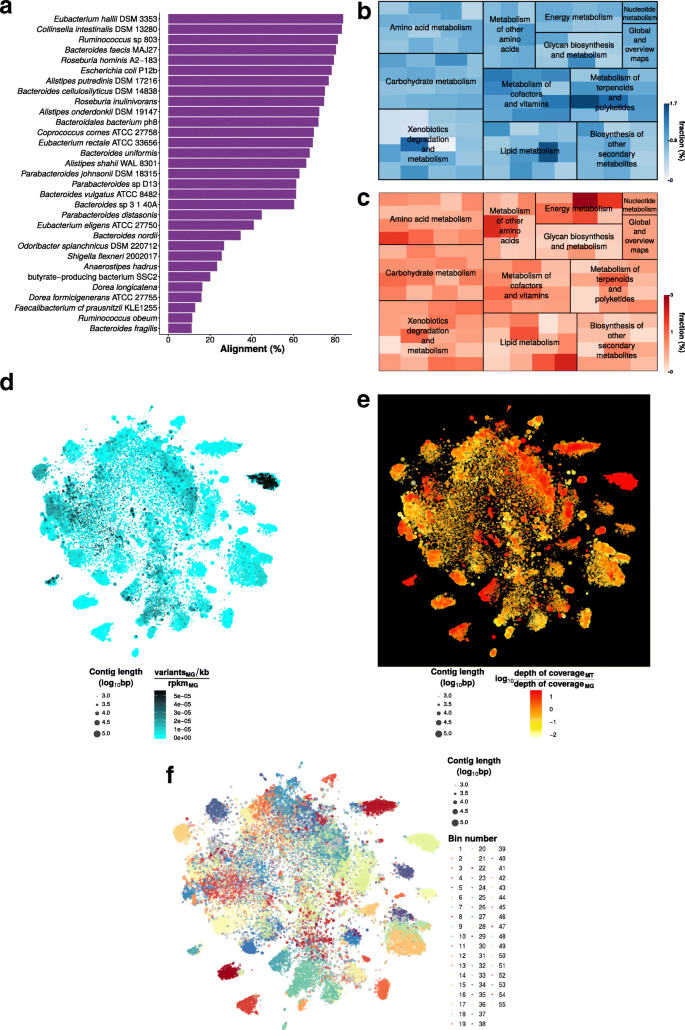
IMP: a pipeline for reproducible reference-independent integrated metagenomic and metatranscriptomic analyses | Genome Biology | Full Text

COVID-19 Disease Map, a computational knowledge repository of SARS-CoV-2 virus-host interaction mechanisms | bioRxiv
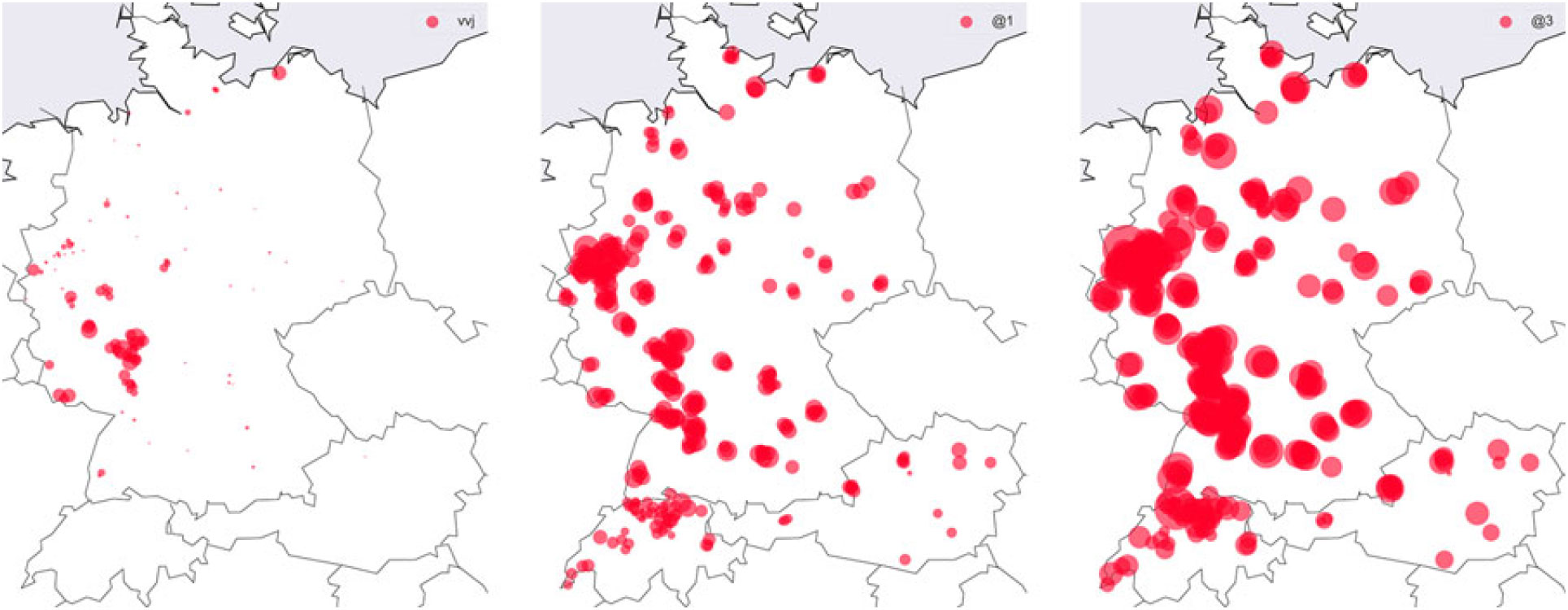
Lörres, Möppes, and the Swiss. (Re)Discovering regional patterns in anonymous social media data | Journal of Linguistic Geography | Cambridge Core

COVID-19 Disease Map, a computational knowledge repository of SARS-CoV-2 virus-host interaction mechanisms | bioRxiv

COVID-19 Disease Map, a computational knowledge repository of SARS-CoV-2 virus-host interaction mechanisms | bioRxiv
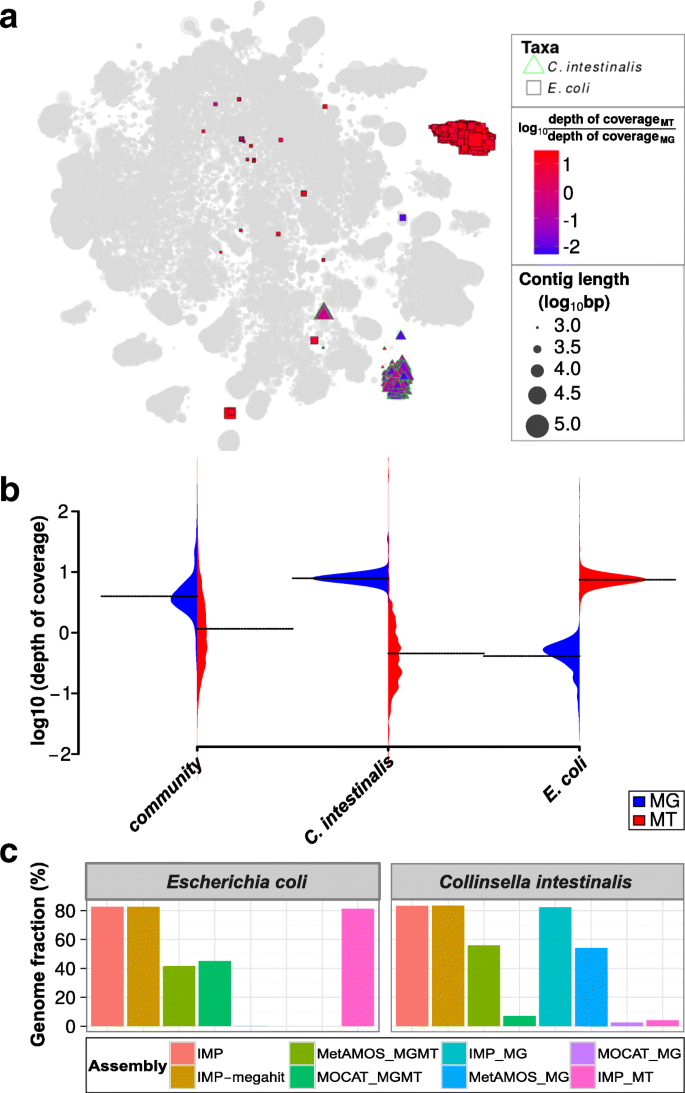
IMP: a pipeline for reproducible reference-independent integrated metagenomic and metatranscriptomic analyses | Genome Biology | Full Text
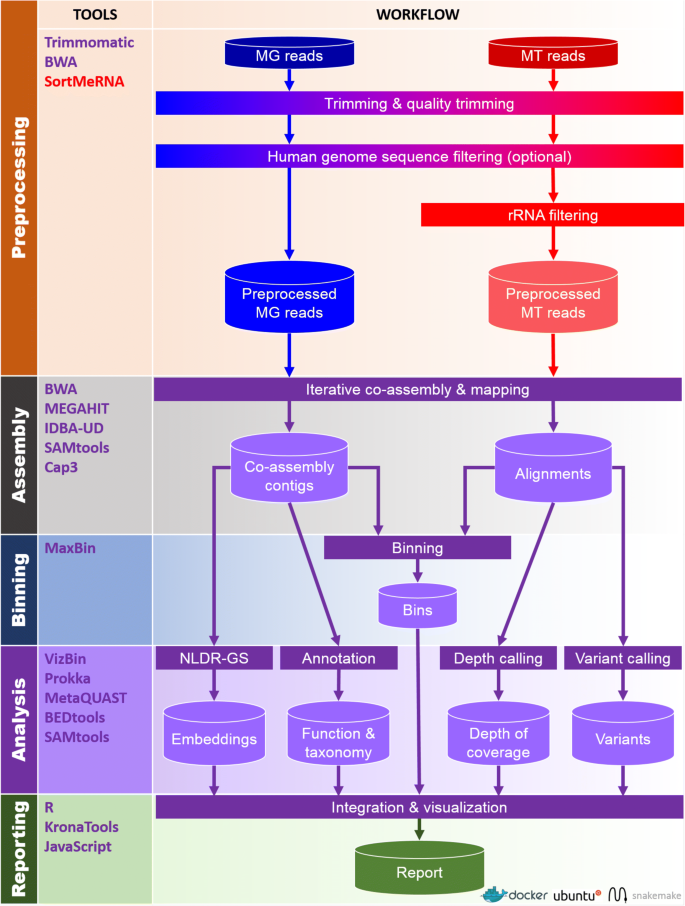
IMP: a pipeline for reproducible reference-independent integrated metagenomic and metatranscriptomic analyses | Genome Biology | Full Text

Genome-wide mega-analysis identifies 16 loci and highlights diverse biological mechanisms in the common epilepsies | Nature Communications

COVID-19 Disease Map, a computational knowledge repository of SARS-CoV-2 virus-host interaction mechanisms | bioRxiv

Modeling seizures in the Human Phenotype Ontology according to contemporary ILAE concepts makes big phenotypic data tractable - Lewis‐Smith - 2021 - Epilepsia - Wiley Online Library
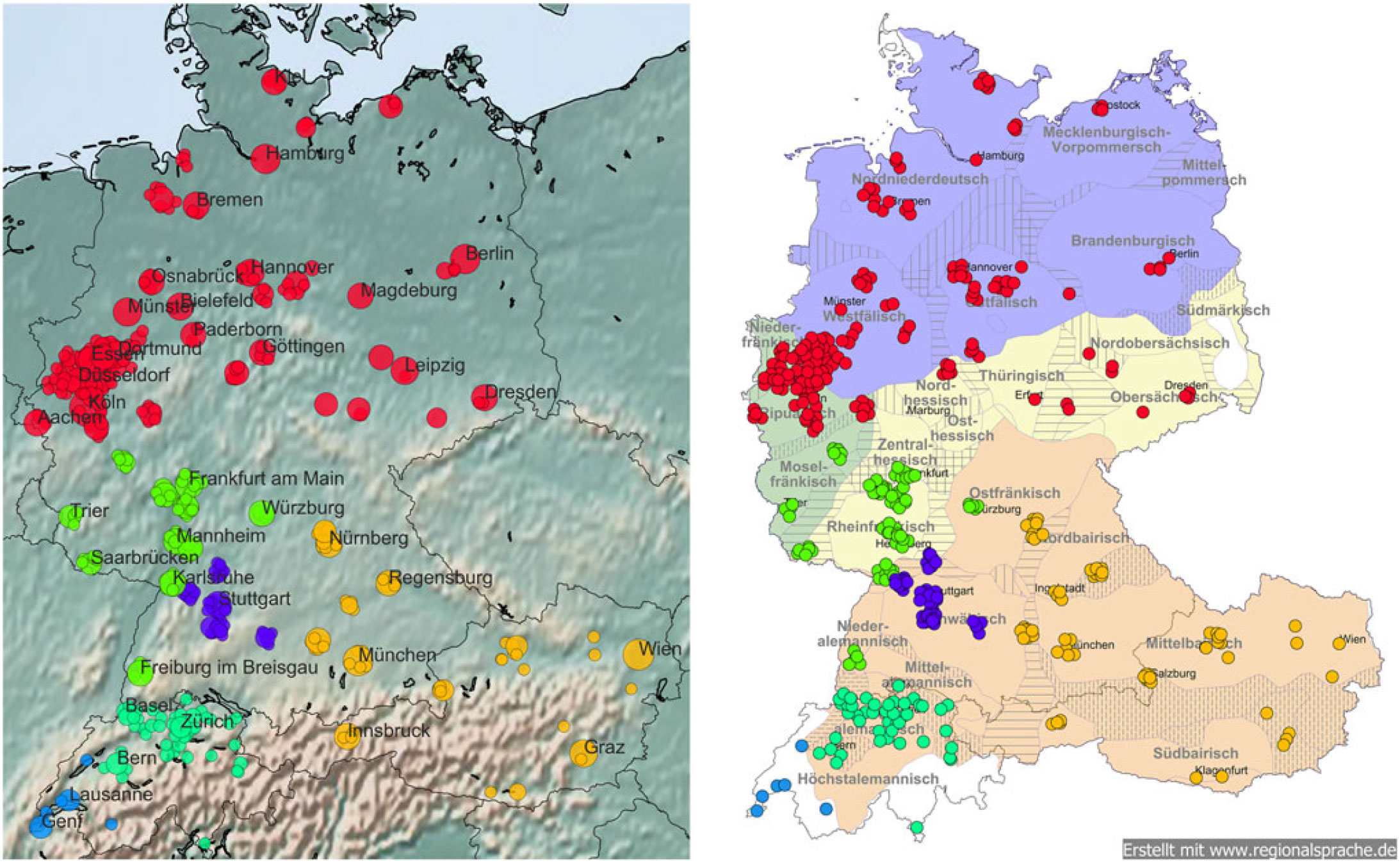
Lörres, Möppes, and the Swiss. (Re)Discovering regional patterns in anonymous social media data | Journal of Linguistic Geography | Cambridge Core

Modeling seizures in the Human Phenotype Ontology according to contemporary ILAE concepts makes big phenotypic data tractable - Lewis‐Smith - 2021 - Epilepsia - Wiley Online Library
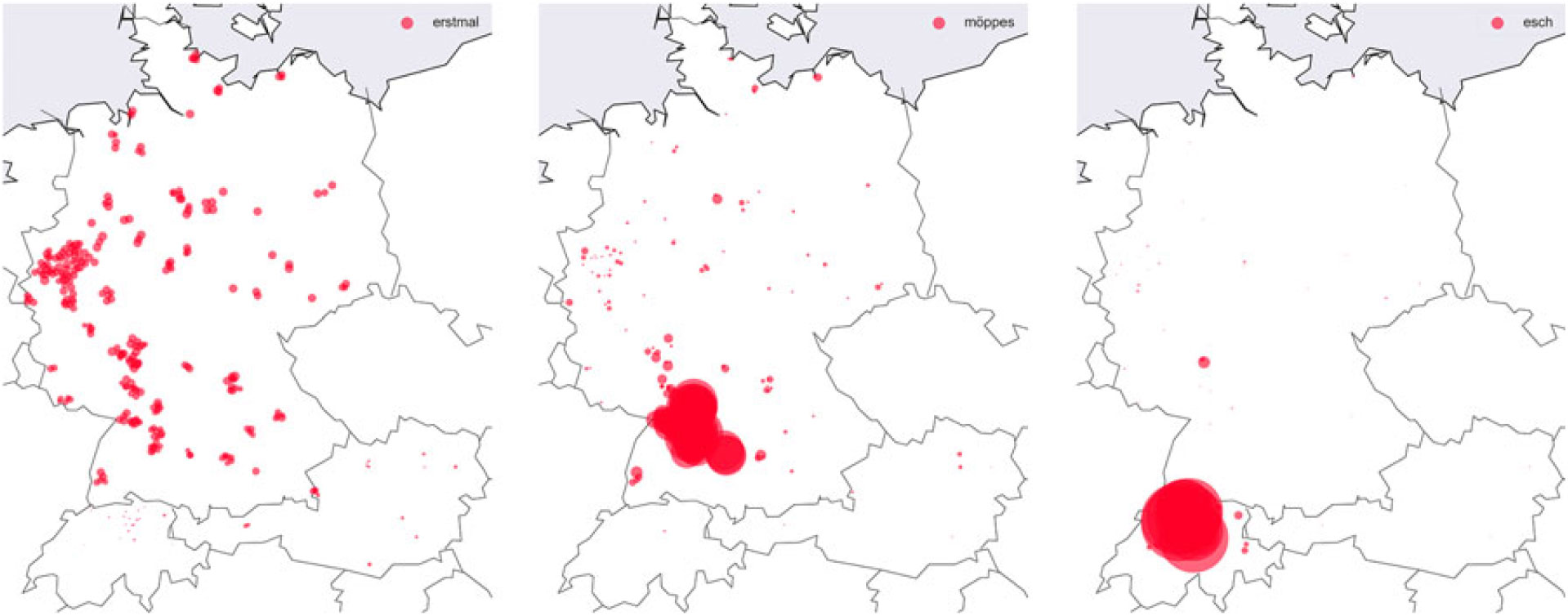
Lörres, Möppes, and the Swiss. (Re)Discovering regional patterns in anonymous social media data | Journal of Linguistic Geography | Cambridge Core